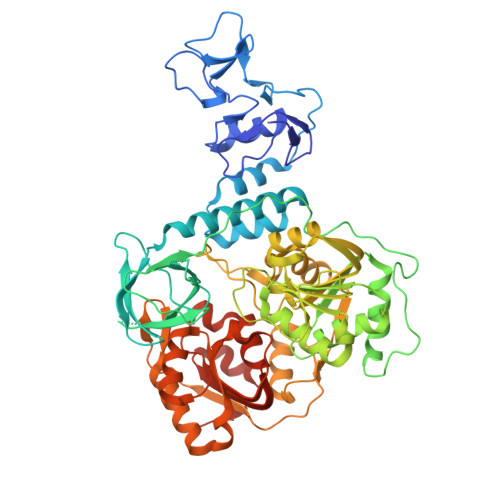
Nucleotide-bound crystal structures of the SARS-CoV-2 helicase NSP13.
Kloskowski, P., Neumann, P., Berndt, A., Ficner, R.(2025) Acta Crystallogr F Struct Biol Commun
- PubMed: 40638074
- DOI: https://doi.org/10.1107/S2053230X25005266
- Primary Citation of Related Structures:
9I53 - PubMed Abstract:
Nucleotide-bound crystal structures of SARS-CoV-2 NSP13 in ADP- and ATP-bound states were resolved to 1.8 and 1.9 Å, respectively. The ADP-bound model captures a state immediately following ATP hydrolysis, with both ADP and orthophosphate still present in the active site. Further comparative analysis revealed that crystal packing influences NSP13 by stabilizing the nucleotide-binding site, underscoring the importance of accounting for these effects in structure-based drug design targeting NSP13.
Organizational Affiliation:
Department of Molecular Structural Biology, Institute of Microbiology and Genetics, Göttingen Center of Molecular Biosciences (GZMB), University of Göttingen, Justus-von-Liebig-Weg 11, 37077 Göttingen, Germany.